Metabolic Networks¶
Metabolic networks have four columns. The first declares a unique name for the enzyme or enzymatic complex; the second declares a unique name for the reaction; the third column lists using comma unique names for substrates; and the last row list using comma unique names for products. To declare metabolites located at the periplasm or extracellular compartments, the user should employ the prefix “PER-” and “EX-”, respectively. Use spontaneous for non-enzymatic reactions.
Examples:
GENE REACTION SUBSTRATES PRODUCTS
spontaneous LACTOSE-MUTAROTATION alpha-lactose beta-lactose
spontaneous GALACTOSE-MUTAROTATION alpha-GALACTOSE beta-GALACTOSE
spontaneous GLUCOSE-MUTAROTATION alpha-glucose beta-glucose
LACY-MONOMER TRANS-RXN-24 PER-PROTON, PER-alpha-lactose PROTON, alpha-lactose
LACY-MONOMER TRANS-RXN-24-beta PER-PROTON, PER-beta-lactose PROTON, beta-lactose
LACY-MONOMER TRANS-RXN-94 PER-PROTON, PER-MELIBIOSE PROTON, MELIBIOSE
LACY-MONOMER RXN0-7215 PER-PROTON, PER-CPD-3561 PROTON, CPD-3561
LACY-MONOMER RXN0-7217 PER-PROTON, PER-CPD-3785 PROTON, CPD-3785
LACY-MONOMER RXN-17755 PER-PROTON, PER-CPD-3801 PROTON, CPD-3801
BETAGALACTOSID-CPLX BETAGALACTOSID-RXN beta-lactose, WATER beta-GALACTOSE, beta-glucose
BETAGALACTOSID-CPLX BETAGALACTOSID-RXN-alpha alpha-lactose, WATER alpha-GALACTOSE, alpha-glucose
BETAGALACTOSID-CPLX RXN0-5363 alpha-lactose alpha-ALLOLACTOSE
BETAGALACTOSID-CPLX RXN0-5363-beta beta-lactose beta-ALLOLACTOSE
BETAGALACTOSID-CPLX ALLOLACTOSE-DEG-alpha alpha-ALLOLACTOSE alpha-GALACTOSE, alpha-glucose
BETAGALACTOSID-CPLX ALLOLACTOSE-DEG-beta beta-ALLOLACTOSE beta-GALACTOSE, beta-glucose
BETAGALACTOSID-CPLX RXN-17726 CPD-3561, WATER beta-GALACTOSE, Fructofuranose
BETAGALACTOSID-CPLX RXN0-7219 CPD-3785, WATER beta-GALACTOSE, D-ARABINOSE
GALACTOACETYLTRAN-CPLX GALACTOACETYLTRAN-RXN-galactose beta-GALACTOSE, ACETYL-COA 6-Acetyl-beta-D-Galactose, CO-A
OR
GENE REACTION SUBSTRATES PRODUCTS
spontaneous LACTOSE-MUTAROTATION alpha-lactose beta-lactose
spontaneous GALACTOSE-MUTAROTATION alpha-GALACTOSE beta-GALACTOSE
spontaneous GLUCOSE-MUTAROTATION alpha-glucose beta-glucose
lacY TRANS-RXN-24 PER-PROTON, PER-alpha-lactose PROTON, alpha-lactose
lacY TRANS-RXN-24-beta PER-PROTON, PER-beta-lactose PROTON, beta-lactose
lacY TRANS-RXN-94 PER-PROTON, PER-MELIBIOSE PROTON, MELIBIOSE
lacY RXN0-7215 PER-PROTON, PER-CPD-3561 PROTON, CPD-3561
lacY RXN0-7217 PER-PROTON, PER-CPD-3785 PROTON, CPD-3785
lacY RXN-17755 PER-PROTON, PER-CPD-3801 PROTON, CPD-3801
[lacZ,lacZ,lacZ,lacZ] BETAGALACTOSID-RXN beta-lactose, WATER beta-GALACTOSE, beta-glucose
[lacZ,lacZ,lacZ,lacZ] BETAGALACTOSID-RXN-alpha alpha-lactose, WATER alpha-GALACTOSE, alpha-glucose
[lacZ,lacZ,lacZ,lacZ] RXN0-5363 alpha-lactose alpha-ALLOLACTOSE
[lacZ,lacZ,lacZ,lacZ] RXN0-5363-beta beta-lactose beta-ALLOLACTOSE
[lacZ,lacZ,lacZ,lacZ] ALLOLACTOSE-DEG-alpha alpha-ALLOLACTOSE, WATER alpha-GALACTOSE, alpha-glucose
[lacZ,lacZ,lacZ,lacZ] ALLOLACTOSE-DEG-beta beta-ALLOLACTOSE, WATER beta-GALACTOSE, beta-glucose
[lacZ,lacZ,lacZ,lacZ] RXN-17726 CPD-3561, WATER beta-GALACTOSE, Fructofuranose
[lacZ,lacZ,lacZ,lacZ] RXN0-7219 CPD-3785, WATER beta-GALACTOSE, D-ARABINOSE
[lacA,lacA,lacA] GALACTOACETYLTRAN-RXN-galactose beta-GALACTOSE, ACETYL-COA 6-Acetyl-beta-D-Galactose, CO-A
OR
GENE REACTION SUBSTRATES PRODUCTS
spontaneous LACTOSE-MUTAROTATION alpha-lactose beta-lactose
spontaneous GALACTOSE-MUTAROTATION alpha-GALACTOSE beta-GALACTOSE
spontaneous GLUCOSE-MUTAROTATION alpha-glucose beta-glucose
lacY TRANS-RXN-24 PER-PROTON, PER-alpha-lactose PROTON, alpha-lactose
lacY TRANS-RXN-24-beta PER-PROTON, PER-beta-lactose PROTON, beta-lactose
lacY TRANS-RXN-94 PER-PROTON, PER-MELIBIOSE PROTON, MELIBIOSE
lacY RXN0-7215 PER-PROTON, PER-CPD-3561 PROTON, CPD-3561
lacY RXN0-7217 PER-PROTON, PER-CPD-3785 PROTON, CPD-3785
lacY RXN-17755 PER-PROTON, PER-CPD-3801 PROTON, CPD-3801
lacZ BETAGALACTOSID-RXN beta-lactose, WATER beta-GALACTOSE, beta-glucose
lacZ BETAGALACTOSID-RXN-alpha alpha-lactose, WATER alpha-GALACTOSE, alpha-glucose
lacZ RXN0-5363 alpha-lactose alpha-ALLOLACTOSE
lacZ RXN0-5363-beta beta-lactose beta-ALLOLACTOSE
lacZ ALLOLACTOSE-DEG-alpha alpha-ALLOLACTOSE, WATER alpha-GALACTOSE, alpha-glucose
lacZ ALLOLACTOSE-DEG-beta beta-ALLOLACTOSE, WATER beta-GALACTOSE, beta-glucose
lacZ RXN-17726 CPD-3561, WATER beta-GALACTOSE, Fructofuranose
lacZ RXN0-7219 CPD-3785, WATER beta-GALACTOSE, D-ARABINOSE
lacA GALACTOACETYLTRAN-RXN-galactose beta-GALACTOSE, ACETYL-COA 6-Acetyl-beta-D-Galactose, CO-A
Note
Visualization in Cytoscape. Transform the four-columns file into a two-columns file with the helper script “Expand metabolic network.ipynb”, paste the results in a new file, and import the network into Cytoscape. Colors and arrows remains to the user for customization.
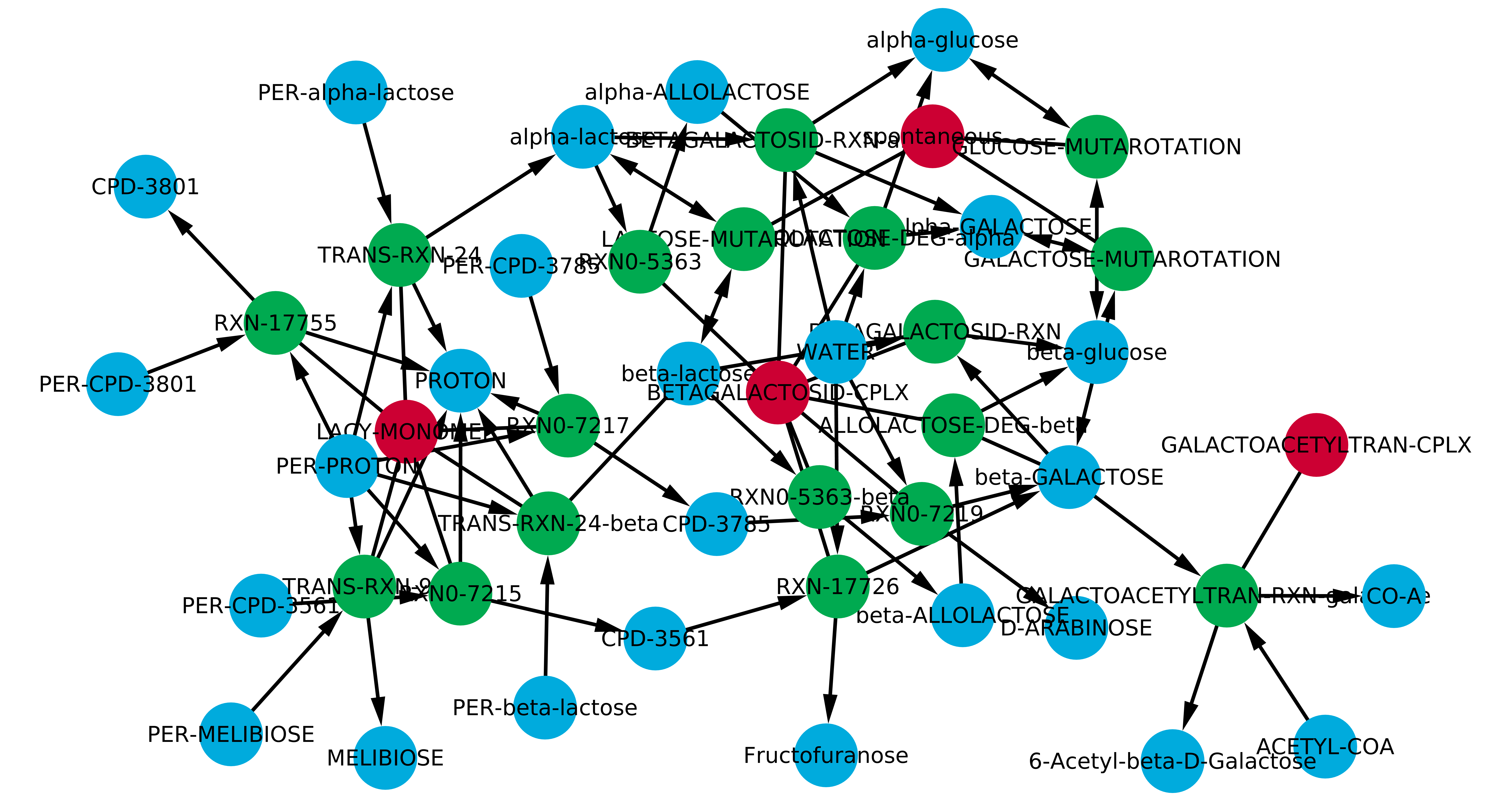
Finally, execute the “Rules from metabolic network.ipynb” to obtain the Rules to model the defined metabolic network. The complete rule-based model can be found in the lactose folder from the Network Biology Lab GitHub repository here.
Rule('LACTOSE_MUTAROTATION',
met(name = 'alpha_lactose', loc = 'cyt') |
met(name = 'beta_lactose', loc = 'cyt'),
Parameter('fwd_LACTOSE_MUTAROTATION', 1),
Parameter('rvs_LACTOSE_MUTAROTATION', 1))
Rule('GALACTOSE_MUTAROTATION',
met(name = 'alpha_GALACTOSE', loc = 'cyt') |
met(name = 'beta_GALACTOSE', loc = 'cyt'),
Parameter('fwd_GALACTOSE_MUTAROTATION', 1),
Parameter('rvs_GALACTOSE_MUTAROTATION', 1))
Rule('GLUCOSE_MUTAROTATION',
met(name = 'alpha_glucose', loc = 'cyt') |
met(name = 'beta_glucose', loc = 'cyt'),
Parameter('fwd_GLUCOSE_MUTAROTATION', 1),
Parameter('rvs_GLUCOSE_MUTAROTATION', 1))
Rule('TRANS_RXN_24',
prot(name = 'LACY_MONOMER') +
met(name = 'PROTON', loc = 'per') +
met(name = 'alpha_lactose', loc = 'per') |
prot(name = 'LACY_MONOMER') +
met(name = 'PROTON', loc = 'cyt') +
met(name = 'alpha_lactose', loc = 'cyt'),
Parameter('fwd_TRANS_RXN_24', 1),
Parameter('rvs_TRANS_RXN_24', 1))
Rule('TRANS_RXN_24_beta',
prot(name = 'LACY_MONOMER') +
met(name = 'PROTON', loc = 'per') +
met(name = 'beta_lactose', loc = 'per') |
prot(name = 'LACY_MONOMER') +
met(name = 'PROTON', loc = 'cyt') +
met(name = 'beta_lactose', loc = 'cyt'),
Parameter('fwd_TRANS_RXN_24_beta', 1),
Parameter('rvs_TRANS_RXN_24_beta', 1))
Rule('TRANS_RXN_94',
prot(name = 'LACY_MONOMER') +
met(name = 'PROTON', loc = 'per') +
met(name = 'MELIBIOSE', loc = 'per') |
prot(name = 'LACY_MONOMER') +
met(name = 'PROTON', loc = 'cyt') +
met(name = 'MELIBIOSE', loc = 'cyt'),
Parameter('fwd_TRANS_RXN_94', 1),
Parameter('rvs_TRANS_RXN_94', 1))
Rule('RXN0_7215', prot(name = 'LACY_MONOMER') +
met(name = 'PROTON', loc = 'per') +
met(name = 'CPD_3561', loc = 'per') |
prot(name = 'LACY_MONOMER') +
met(name = 'PROTON', loc = 'cyt') +
met(name = 'CPD_3561', loc = 'cyt'),
Parameter('fwd_RXN0_7215', 1),
Parameter('rvs_RXN0_7215', 1))
Rule('RXN0_7217', prot(name = 'LACY_MONOMER') +
met(name = 'PROTON', loc = 'per') +
met(name = 'CPD_3785', loc = 'per') |
prot(name = 'LACY_MONOMER') +
met(name = 'PROTON', loc = 'cyt') +
met(name = 'CPD_3785', loc = 'cyt'),
Parameter('fwd_RXN0_7217', 1),
Parameter('rvs_RXN0_7217', 1))
Rule('RXN_17755', prot(name = 'LACY_MONOMER') +
met(name = 'PROTON', loc = 'per') +
met(name = 'CPD_3801', loc = 'per') |
prot(name = 'LACY_MONOMER') +
met(name = 'PROTON', loc = 'cyt') +
met(name = 'CPD_3801', loc = 'cyt'),
Parameter('fwd_RXN_17755', 1),
Parameter('rvs_RXN_17755', 1))
Rule('BETAGALACTOSID_RXN',
cplx(name = 'BETAGALACTOSID_CPLX') +
met(name = 'beta_lactose', loc = 'cyt') +
met(name = 'WATER', loc = 'cyt') |
cplx(name = 'BETAGALACTOSID_CPLX') +
met(name = 'beta_GALACTOSE', loc = 'cyt') +
met(name = 'beta_glucose', loc = 'cyt'),
Parameter('fwd_BETAGALACTOSID_RXN', 1),
Parameter('rvs_BETAGALACTOSID_RXN', 1))
Rule('BETAGALACTOSID_RXN_alpha',
cplx(name = 'BETAGALACTOSID_CPLX') +
met(name = 'alpha_lactose', loc = 'cyt') +
met(name = 'WATER', loc = 'cyt') |
cplx(name = 'BETAGALACTOSID_CPLX') +
met(name = 'alpha_GALACTOSE', loc = 'cyt') +
met(name = 'alpha_glucose', loc = 'cyt'),
Parameter('fwd_BETAGALACTOSID_RXN_alpha', 1),
Parameter('rvs_BETAGALACTOSID_RXN_alpha', 1))
Rule('RXN0_5363',
cplx(name = 'BETAGALACTOSID_CPLX') +
met(name = 'alpha_lactose', loc = 'cyt') |
cplx(name = 'BETAGALACTOSID_CPLX') +
met(name = 'alpha_ALLOLACTOSE', loc = 'cyt'),
Parameter('fwd_RXN0_5363', 1),
Parameter('rvs_RXN0_5363', 1))
Rule('RXN0_5363_beta',
cplx(name = 'BETAGALACTOSID_CPLX') +
met(name = 'beta_lactose', loc = 'cyt') |
cplx(name = 'BETAGALACTOSID_CPLX') +
met(name = 'beta_ALLOLACTOSE', loc = 'cyt'),
Parameter('fwd_RXN0_5363_beta', 1),
Parameter('rvs_RXN0_5363_beta', 1))
Rule('ALLOLACTOSE_DEG_alpha',
cplx(name = 'BETAGALACTOSID_CPLX') +
met(name = 'alpha_ALLOLACTOSE', loc = 'cyt') |
cplx(name = 'BETAGALACTOSID_CPLX') +
met(name = 'alpha_GALACTOSE', loc = 'cyt'),
Parameter('fwd_ALLOLACTOSE_DEG_alpha', 1),
Parameter('rvs_ALLOLACTOSE_DEG_alpha', 1))
Rule('ALLOLACTOSE_DEG_beta',
cplx(name = 'BETAGALACTOSID_CPLX') +
met(name = 'beta_ALLOLACTOSE', loc = 'cyt') |
cplx(name = 'BETAGALACTOSID_CPLX') +
met(name = 'beta_GALACTOSE', loc = 'cyt'),
Parameter('fwd_ALLOLACTOSE_DEG_beta', 1),
Parameter('rvs_ALLOLACTOSE_DEG_beta', 1))
Rule('RXN_17726',
cplx(name = 'BETAGALACTOSID_CPLX') +
met(name = 'CPD_3561', loc = 'cyt') +
met(name = 'WATER', loc = 'cyt') |
cplx(name = 'BETAGALACTOSID_CPLX') +
met(name = 'beta_GALACTOSE', loc = 'cyt') +
met(name = 'Fructofuranose', loc = 'cyt'),
Parameter('fwd_RXN_17726', 1),
Parameter('rvs_RXN_17726', 1))
Rule('RXN0_7219',
cplx(name = 'BETAGALACTOSID_CPLX') +
met(name = 'CPD_3785', loc = 'cyt') +
met(name = 'WATER', loc = 'cyt') |
cplx(name = 'BETAGALACTOSID_CPLX') +
met(name = 'beta_GALACTOSE', loc = 'cyt') +
met(name = 'D_ARABINOSE', loc = 'cyt'),
Parameter('fwd_RXN0_7219', 1),
Parameter('rvs_RXN0_7219', 1))
Rule('GALACTOACETYLTRAN_RXN_galactose',
cplx(name = 'GALACTOACETYLTRAN_CPLX') +
met(name = 'beta_GALACTOSE', loc = 'cyt') +
met(name = 'ACETYL_COA', loc = 'cyt') |
cplx(name = 'GALACTOACETYLTRAN_CPLX') +
met(name = '_6_Acetyl_beta_D_Galactose', loc = 'cyt') +
met(name = 'CO_A', loc = 'cyt'),
Parameter('fwd_GALACTOACETYLTRAN_RXN_galactose', 1),
Parameter('rvs_GALACTOACETYLTRAN_RXN_galactose', 1))
Note
Reversibility of Rules. Atlas writes reversible Rules for each
reaction declared in the network file. The Parameter('rvs_RuleName', 1))
must be set to zero to define an irreversible reaction.
Note
Uniqueness of Rule names. Atlas will write Rules for unique metabolic reactions. Identical names will be reported for further curation.
Note
Simulation. The model can be simulated only with the instantiation of
Monomers and Initials (More here).
Run Monomer+Initials+Observables from metabolic network.ipynb to obtain
automatically the necessary Monomers and Initials (including
proteins and enzymatic complexes).
Plotting. The model can be observed only with the instantation of
Observables (More here).
Run Monomer+Initials+Observables from metabolic network.ipynb to obtain
automatically the all possible Observables for metabolites.